 I am incredibly honored to announce my election to the Board of Trustees of the New York Genealogical & Biographical Society! The NYG&B is the largest and oldest genealogical society in New York State, and the second oldest genealogical society in the nation. As a lifelong genealogist with New York roots dating back almost 250 years, joining the NYG&B is a dream come true for me.
I am incredibly honored to announce my election to the Board of Trustees of the New York Genealogical & Biographical Society! The NYG&B is the largest and oldest genealogical society in New York State, and the second oldest genealogical society in the nation. As a lifelong genealogist with New York roots dating back almost 250 years, joining the NYG&B is a dream come true for me.
Over the past decade, DNA has become a powerful tool for genealogical research. As a member of the NYG&B’s Board of Trustees, I hope to be able to help bridge the (ever-closing) gap between traditional genealogy and genetic genealogy, and help both members and non-members understand and incorporate DNA into their family histories.
The board represents an incredible group of people dedicated to helping people discovery their family histories, and I am so grateful to be able to join them. The full list is below.

 If you’re serious about genetic genealogy, you’ve heard of
If you’re serious about genetic genealogy, you’ve heard of 
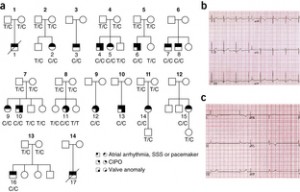 In 2008, I wrote about the case of Mr. and Mrs. George Fry, who are believed to have brought a particularly negative mutation with them to the New World from Europe in 1630 (“
In 2008, I wrote about the case of Mr. and Mrs. George Fry, who are believed to have brought a particularly negative mutation with them to the New World from Europe in 1630 (“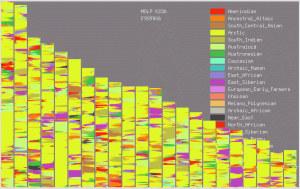
 I’ve written before about a poster presented by AncestryDNA at the American Society of Human Genetics 2013 annual meeting, entitled “
I’ve written before about a poster presented by AncestryDNA at the American Society of Human Genetics 2013 annual meeting, entitled “ This week in Vox, health reporter Julia Belluz (
This week in Vox, health reporter Julia Belluz ( The latest announcement by the newly founded
The latest announcement by the newly founded