Are you interested in genealogy? Are you interested in DNA? Are you interested in Alaska?
You didn’t really say “no” to any of those, did you??
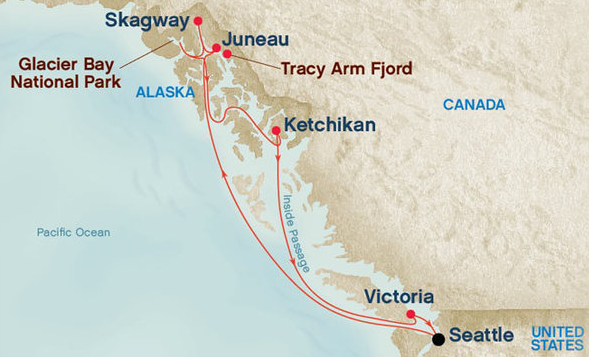
Heritage Books, a leader in the world of genealogy publication for four decades, is again hosting the Annual Heritage Books Genealogy Conference and Cruise! This year’s cruise on Princess Cruise Lines is in Alaska! The cruise departs from Seattle, Washington on September 17, 2017 and arrives back one week later on September 24, 2017.
And just look at this itinerary!
- Day 1 – Sep 17, 2017 (Sunday) – Seattle — Departs at 4:00 PM (First one-on-one session)
- Day 2 – At Sea — Genealogy Sessions (Sessions 1-6)
- Day 3 – Ketchikan, Alaska — Arrives at 6:30 AM and departs 3:00 PM (Second one-on-one session)
- Day 4 – Juneau, Alaska — Arrives at 12:30 PM and departs 10:00 PM (Sessions 7-8)
- Day 5 – Skagway, Alaska — Arrives at 6:00 AM and departs at 5:00 PM (Third one-on-one session)
- Day 6 – At Sea — Genealogy Sessions (Sessions 9-14)
- Day 7 – Victoria, British Columbia — Arrives at 7:00 PM and departs 11:59 PM (Sessions 15-20)
- Day 8 – Sep 24, 2017 (Sunday) – Seattle, Washington — Arrives at 7:00 AM
Great Speakers
This year’s cruise has a terrific line-up of speakers, one I am incredibly honored to join.
- Debra Mieszala, CG – Debbie is a BCG trustee who specializes in forensic genealogy, 20th-century research, and the Midwest. She conducts genealogical research for the military to locate relatives of service members missing in past wars.
- Pamela Boyer Sayre, CG, CGL, FUGA – Pam is a coordinator and teacher at SLIG, IGHR, GRIP, and Boston University’s genealogy certificate program; former NGS director of education and publications; and former board member, NGS and FGS.
- Rick Sayre, CG, CGL, FUGA – Rick is a course coordinator and instructor for IGHR, SLIG, and GRIP. Areas of interest include, urban research, maps, government records, law, land, military, and technology.
- Craig R. Scott, MA, CG, FUGA – Craig is the author of The ‘Lost Pensions’: Settled Accounts of the Act of 6 April 1838 (Revised) and Records of the Accounting Officers of the Department of the Treasury, Inventory 14 (Revised). He has authored seventeen books and several articles in the National Genealogical Society Quarterly Family Chronicle, and other genealogical publications. He is the President and CEO of Heritage Books, Inc., a genealogical publishing firm with over 5,300 titles in print.
- Blaine Bettinger, Ph.D., J.D. – I’m an intellectual property attorney by day and a DNA specialist by night. I’m the author of the long-running blog The Genetic Genealogist, and I frequently give presentations and webinars to educate others about the use of DNA to explore their ancestry.
A Special Offer – Book Now Through February 28th, 2017!
I really want you to join me on this cruise; the more the merrier! From today through February 28th, 2017 only, if you sign up for the cruise and send me proof of your booking (also dated between today and February 28th), I will send you a FREE and SIGNED copy of The Family Tree Guide to DNA Testing and Genetic Genealogy!