In the past week there have been so many articles and posts about either genetic genealogy or DTC genetics that I’m writing them up as a summary post rather than individually.
The New York Times Tackles DTC Genetic Testing
An article in yesterday’s New York Times by Jane E. Brody – “Buyer Beware of Home DNA Tests†– argues that DTC genetic testing is fraught with danger (the article and some of Brody’s arguments are summarized by Grace Ibay of Genetics & Health: “Seven Reasons Why Home DNA Tests Are Hypeâ€). The author even lumps in genetic genealogy (which has been around for over 9 years now, hardly a “new industry†that has sprung up “to cash in†on new science):
 Five bioethicists have published a paper in today’s issue of Science –
Five bioethicists have published a paper in today’s issue of Science – 
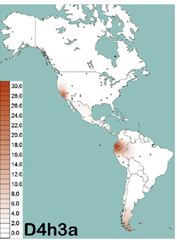 An international team of researchers have concluded that humans entered the Americas from Asia along at least two different paths. By studying two rare mtDNA haplogroups found in Native Americans – D4h3 and X2a – the researchers conclude that D4h3 spread into the Americans along the Pacific coast while X2a entered through the ice-free corridor between the
An international team of researchers have concluded that humans entered the Americas from Asia along at least two different paths. By studying two rare mtDNA haplogroups found in Native Americans – D4h3 and X2a – the researchers conclude that D4h3 spread into the Americans along the Pacific coast while X2a entered through the ice-free corridor between the 