[PLEASE NOTE: The Onion is a satirical site meant for ENTERTAINMENT and social commentary purposes only. The following study is NOT real!]
The Onion, an infamous mock news site has a (surprisingly intelligent) article today entitled “7 Million People Direct Descendants Of Single Smooth-Talking Ancestor” about a “study” that has found that millions of people around the world have a genetic marker that links them to “a single smooth-talking common ancestor.”
Randy Seaver of Genea-Musings brings the article to my attention (thank you Randy!):
The headline screams “7 Million People Direct Descendants of Smooth-Talking Ancestor” — see the article here in the Science and Technology section of The Onion. It sounds right up the genetic genealogy alley, doesn’t it? Megan, Blaine, Emily – why haven’t you written about this guy? Are 7 million descendants not enough?
 An article in the United Arab Emirate newspaper
An article in the United Arab Emirate newspaper 
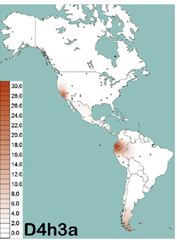 An international team of researchers have concluded that humans entered the Americas from Asia along at least two different paths. By studying two rare mtDNA haplogroups found in Native Americans – D4h3 and X2a – the researchers conclude that D4h3 spread into the Americans along the Pacific coast while X2a entered through the ice-free corridor between the
An international team of researchers have concluded that humans entered the Americas from Asia along at least two different paths. By studying two rare mtDNA haplogroups found in Native Americans – D4h3 and X2a – the researchers conclude that D4h3 spread into the Americans along the Pacific coast while X2a entered through the ice-free corridor between the 



 GenomeWeb Daily News published a story on Friday entitled “
GenomeWeb Daily News published a story on Friday entitled “