If you aren’t already a member of the coolest Facebook group ever, Genetic Genealogy Tips & Techniques, you really should be! We have a friendly and engaging environment, and everyone learns something new every day!
This post is meant to answer a question or issue that is raised almost daily in the group, and that is the issue of small shared DNA segments. Although these small segments are alluring, they are the mythological sirens of the genealogical world!
Small Segments Executive Summary
Here’s a bite-sized summary of the content below:
- Many to most small segments (at least 7 cM and smaller) are FALSE, meaning they are NOT actually shared by the two matches, and therefore do NOT indicate shared ancestry;
- This is supported by a 2014 paper by 23andMe scientists showing that at least 33% of 5 cM phased DNA segments are false-positive (and it’s much worse for unphased segments or segments smaller than 5 cM);
- This is further supported by evidence that anywhere from 20-35% of distant matches at a testing company are not shared with either tested parent;
- This is further supported by evidence that phasing your DNA with two tested parents significantly reduces the number of matches below 10 cM (with proportionally more matches reduced as the segment size gets smaller);
- There is currently no evidence that triangulating segments or finding a paper trail provides a mechanism for distinguishing between false segments and valid segments;
- Since we can’t tell the difference between false small segments and valid small segments, we must avoid these small segments to avoid poisoning our genealogical conclusions with false data; and
- Beware any research or conclusion that uses these small segments without specifically addressing the issues that are known – based on all the scientific research and evidence gathered to date – to surround small segments.
If you’re interested in learning more, keep reading!
Small Segments In Detail
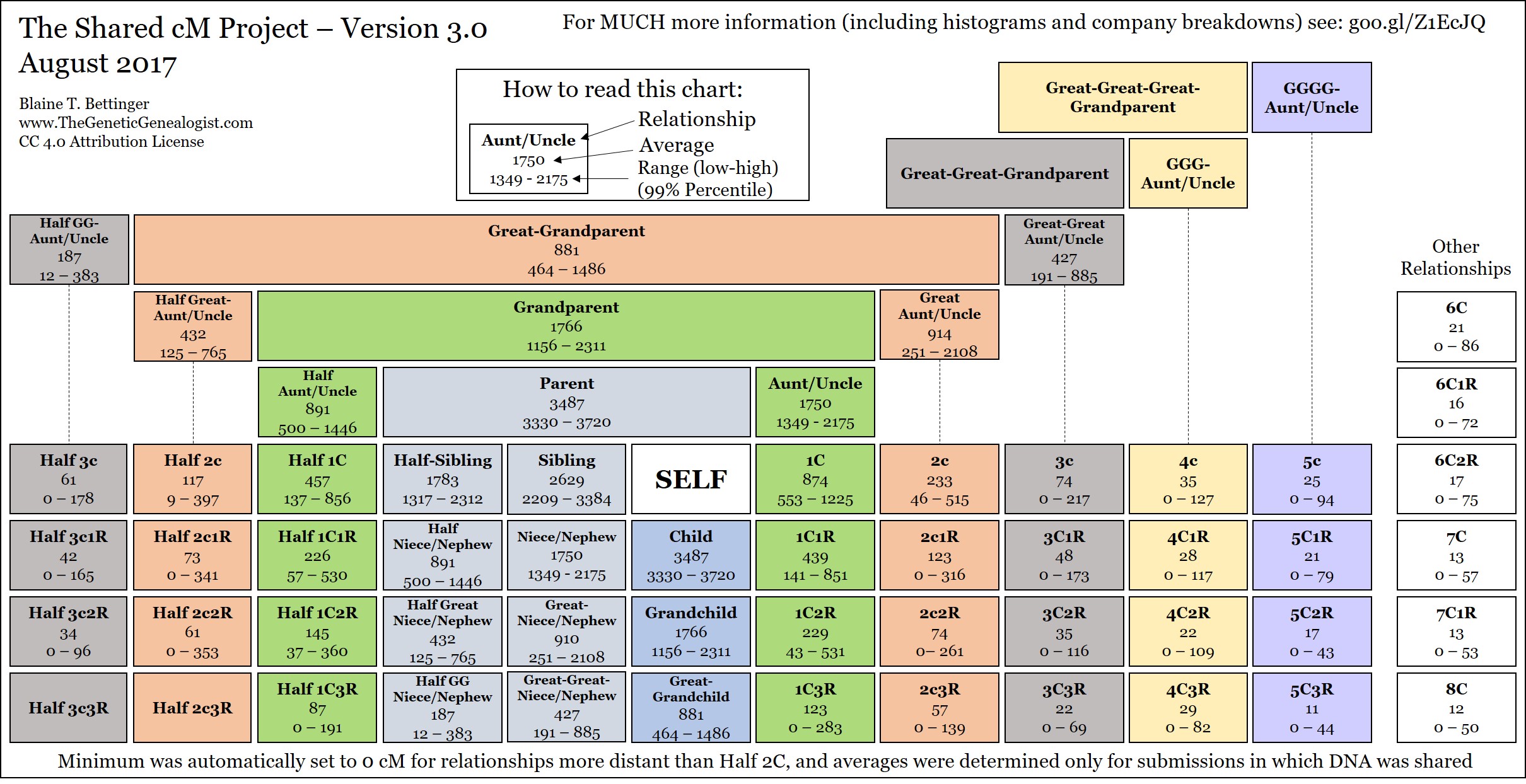
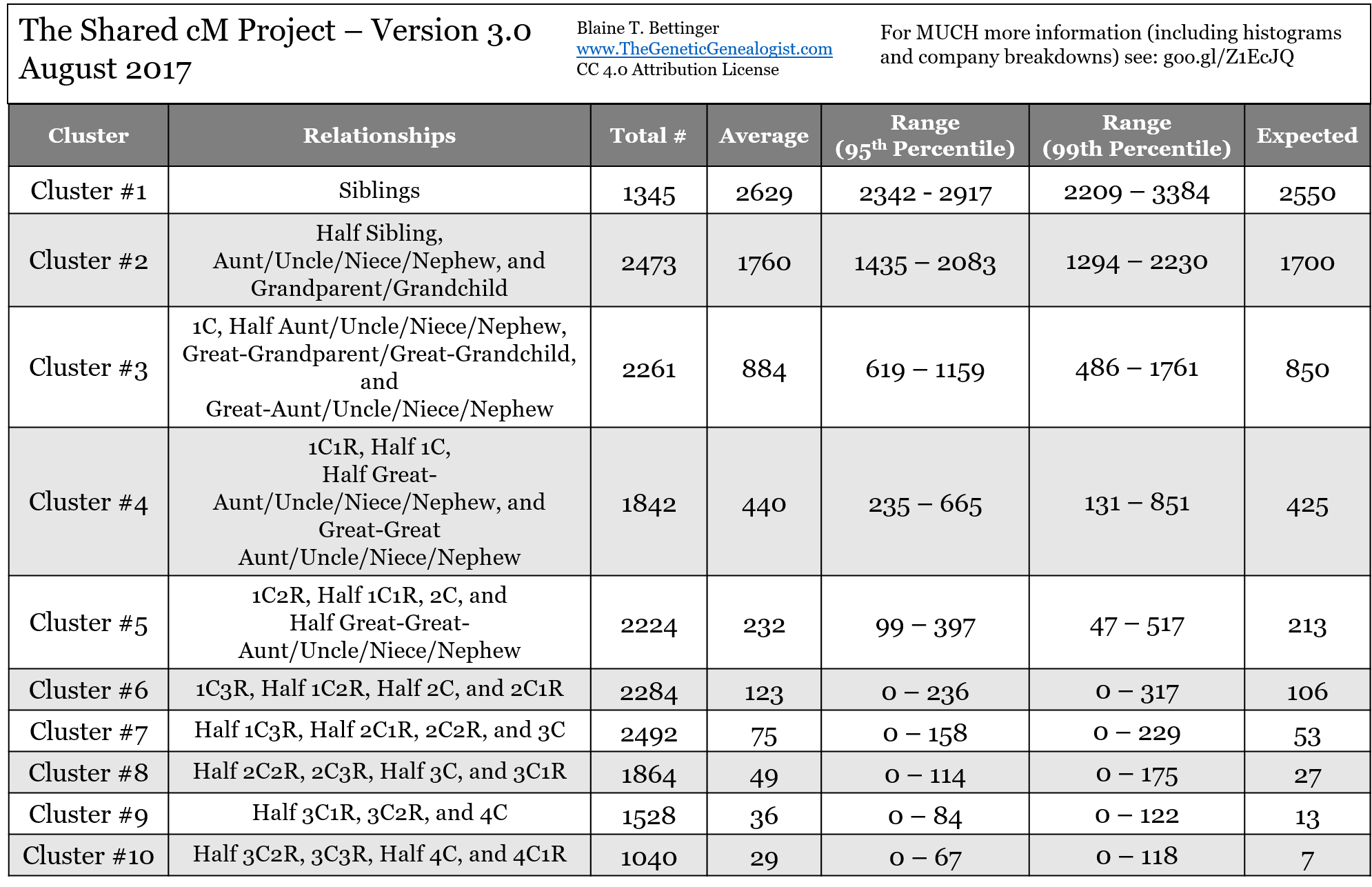
One of the most common questions in the group has to do with small segments. There’s no exact definition of “small” when it comes to small segments, but many of us define them as being a single segment of DNA of 7 cM or smaller. Others use 5 cM or smaller, while others use 10 cM or smaller. Personally, I consider segments of 7 cM or less to be “small,” although when I’m being very conservative I use a definition of 10 cM or smaller. ... Click to read more!