Both 23andMe and deCODEme (using my 23andMe data) have interpreted my SNP results to indicate that I have a greatly increased genetic risk for Type 2 Diabetes. This post interprets the information from both companies and applies some of the primary research that the companies relied upon to predict my risk. Hopefully, this information will be useful to me as I strive to more completely understand my own risk factors, and will be useful to others as an example of using SNP data to potentially understand more about your health.
I. The Genetics
My 23andMe analysis makes it clear that I have an elevated risk for type 2 diabetes:

And, upon clicking upon the link, I receive the following additional information:

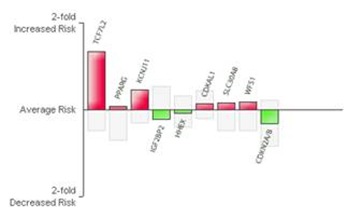
deCODEme, which used my 23andMe data, provides a similar interpretation:

